Determination Of Protein Concentration By Uv Spectrophotometer

So, there I was, hunched over a bubbling beaker, a lab coat that was probably more stained than white, feeling like a mad scientist. My mission? To figure out exactly how much protein was in this mysterious cloudy solution I'd whipped up. You know, the kind of stuff that looks impressive but leaves you wondering if you accidentally created liquid gold or just a really expensive science experiment gone wrong. I’d heard whispers, hushed conversations in the lab about something called a "UV spectrophotometer." It sounded fancy, a bit intimidating, like something you'd find in a James Bond villain's lair. But apparently, it held the key to my protein-counting woes.
Let me tell you, in the world of science, where precision is king (and sometimes queen, let's be inclusive!), knowing the concentration of your precious biomolecules is a pretty big deal. Imagine trying to bake a cake without knowing how much flour you've got. You might end up with a rock-hard brick or a puddle of goo. Proteins are kind of the same. Too little, and your experiment might not work. Too much, and… well, you might get unexpected results, or just waste valuable resources. And that's where our friend, the UV spectrophotometer, waltzes in, ready to save the day.
Think of it like this: you've got a bunch of tiny, invisible protein molecules swimming around in a liquid. How do you count them without a microscope the size of a bus? You can't exactly line them up and tick them off, right? That would be a full-time job, and frankly, I'd rather be doing literally anything else. This is where the magic of light comes in. And not just any light, but ultraviolet (UV) light. Fancy, huh?
The Not-So-Secret Ingredient: Amino Acids
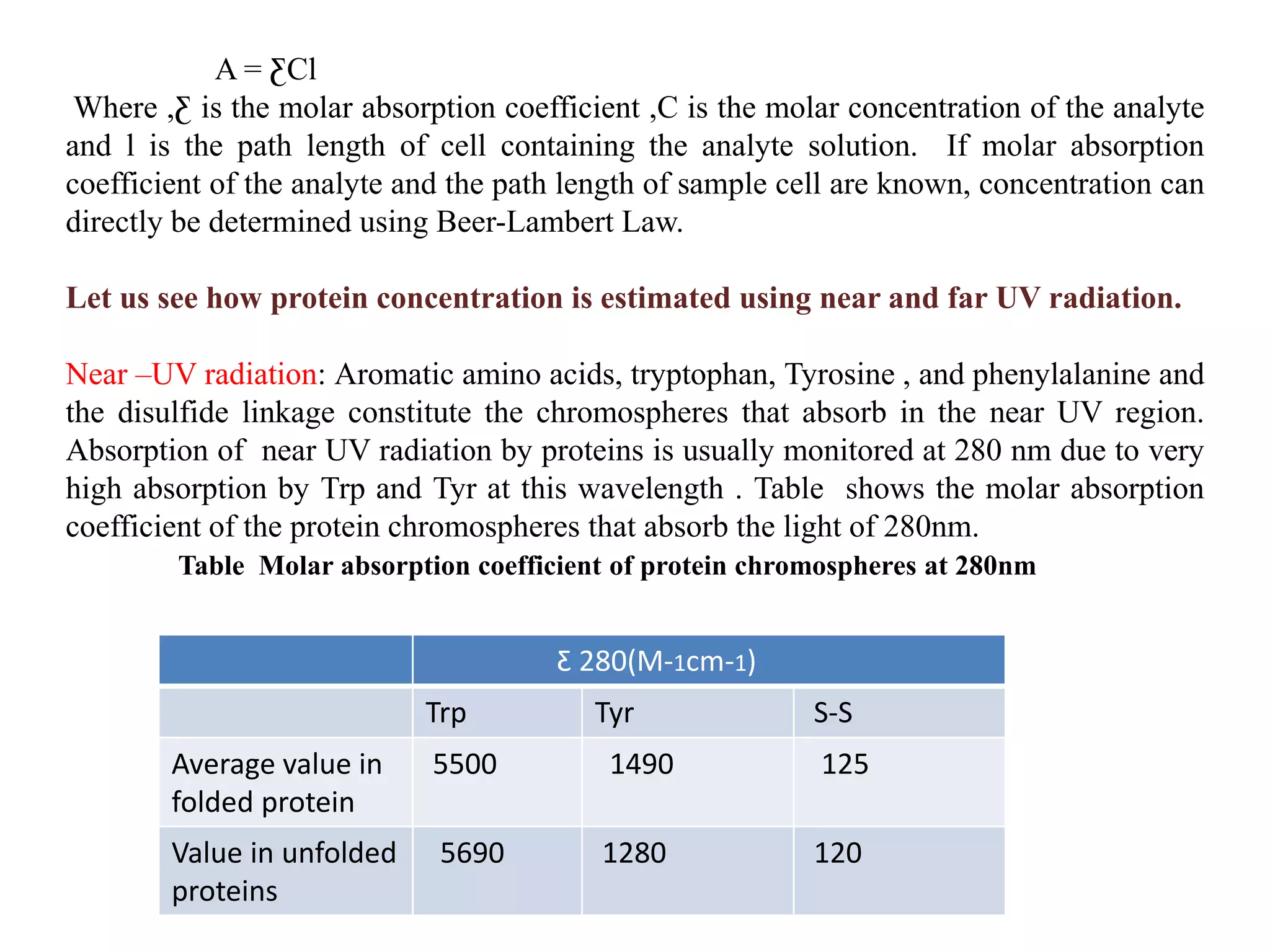
So, why UV light? What’s so special about it when it comes to proteins? Well, it all comes down to the building blocks of proteins: amino acids. Remember those from biology class? They're the little chains that link together to form those big, complex protein structures. Now, some of these amino acids are a bit of a party animal. Specifically, a few of them have these funky little rings in their structure. We're talking about tryptophan, tyrosine, and, to a lesser extent, phenylalanine.
These aromatic rings are like little sponges for UV light. When UV light hits them, they absorb it. It's like they're saying, "Ooh, shiny! Mine!" And the cooler thing is, the amount of UV light they absorb is directly related to how many of these ring-containing amino acids are present. Since most proteins are made up of a mix of amino acids, and the ones with these rings are relatively common (especially tryptophan and tyrosine, which are quite prevalent), we can use this absorption phenomenon as a clever proxy for the total amount of protein. Pretty neat, right? It’s like getting a general headcount of your protein party by looking at how many of your guests are wearing those specific neon party hats.
Enter the UV Spectrophotometer: Your Protein Detective
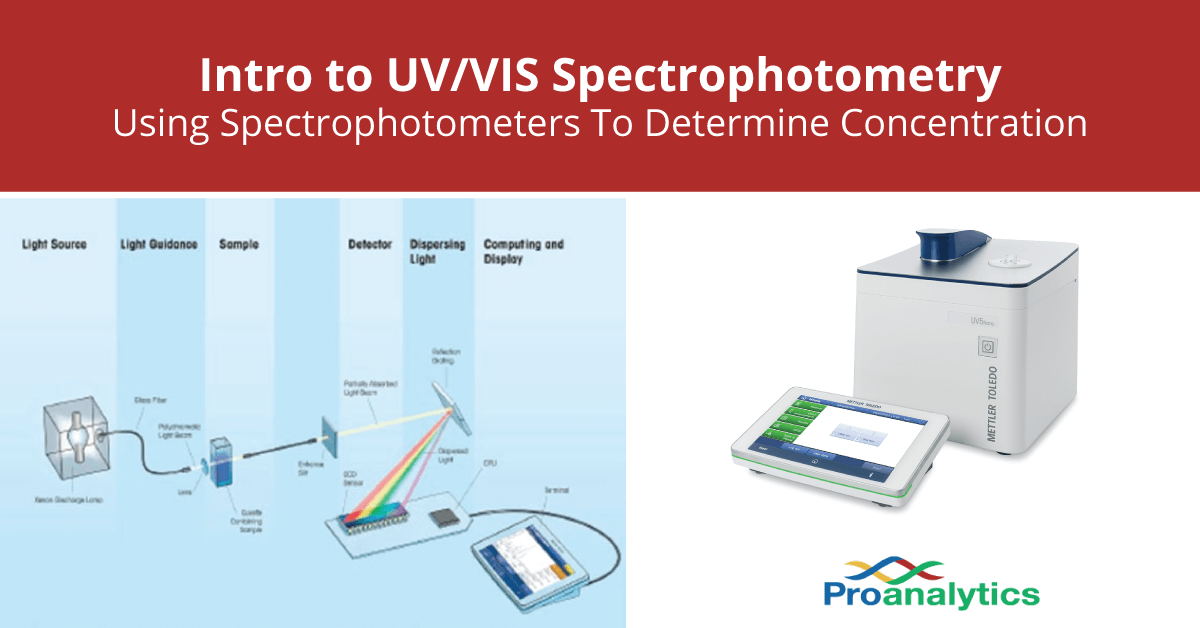
Okay, so we know UV light gets absorbed. But how do we measure that absorption? That's where the spectrophotometer comes in. This contraption is basically a sophisticated light meter. You put your sample in a special little clear box called a cuvette, which is designed not to mess with the light. Then, you shine a beam of UV light through it. The spectrophotometer measures how much light goes in and how much light comes out.
The difference between the light that went in and the light that came out? That's the light that was absorbed by your sample. If your protein solution is clear and barely absorbs any light, you've got a low concentration. If it's like a black hole for UV light, you've got a whole lot of protein in there! It’s like shining a flashlight through a glass of water versus a cup of dark coffee. You’ll notice a big difference in how much light makes it through, right?

The spectrophotometer then spits out a number, usually called absorbance or optical density (OD). This is your raw data. It's a unitless number, which can be a bit confusing at first, but just think of it as a measure of "how dark the sample is to UV light." Higher absorbance means more UV light was soaked up, which means, you guessed it, more protein.
The Wavelength: Picking the Right "Color" of UV Light
Now, you can't just blast your protein sample with any old UV light. Different molecules absorb light at different wavelengths. For proteins, the sweet spot is typically around 280 nanometers (nm). Why 280 nm? Because that's the wavelength where tryptophan and tyrosine really shine (or, more accurately, absorb!). By measuring the absorbance specifically at this wavelength, we’re targeting the protein's absorption and minimizing interference from other things in the solution that might soak up UV light at different wavelengths.
It's like tuning your radio to the exact frequency where your favorite station is broadcasting. You don't want to be on a fuzzy channel, you want the crystal-clear signal. For proteins, 280 nm is our crystal-clear channel. Of course, there are exceptions and other methods, but for a quick, easy, and pretty reliable estimate, 280 nm is the go-to.
The Beer-Lambert Law: The Golden Rule of Absorbance
So, we have absorbance, and we know the wavelength. But how do we get from absorbance to an actual concentration (like milligrams per milliliter, mg/mL)? This is where a fundamental principle in chemistry comes into play: the Beer-Lambert Law. Don't let the name scare you; it's actually quite elegant in its simplicity.
In essence, the Beer-Lambert Law states that the absorbance of a solution is directly proportional to the concentration of the absorbing species and the path length of the light through the solution. Mathematically, it looks like this: A = εbc. Where:
- A is the absorbance (what our spectrophotometer tells us).
- ε (epsilon) is the molar absorptivity, a constant specific to the substance being measured at a particular wavelength. It tells us how strongly a substance absorbs light.
- b is the path length, which is usually the width of our cuvette (typically 1 cm).
- c is the concentration of the absorbing substance.
See? If we know 'A' (from our spectrophotometer), 'b' (the cuvette width), and 'ε' (the absorptivity of our protein), we can rearrange the equation to solve for 'c', the concentration. Ta-da! We've gone from a light-reading to a number that tells us how much protein we actually have.
The Not-So-Perfect World: Factors That Can Mess Things Up
Now, before you go declaring yourself the world's leading protein quantifier, a word of caution. The Beer-Lambert Law, while powerful, works best under ideal conditions. And let's be honest, science is rarely ideal. There are a few things that can throw a wrench in the works:
Interference: Remember how we said proteins absorb UV light because of those amino acid rings? Well, other molecules in your solution might also absorb UV light at 280 nm. Things like nucleic acids (DNA and RNA) are notorious for absorbing strongly at this wavelength. If you have DNA floating around in your protein sample, your absorbance reading will be higher than it should be, leading you to overestimate your protein concentration. It's like trying to count people in a crowd, but some of them are wearing really tall hats, making them look like two people.
Concentration Range: The Beer-Lambert Law is most accurate at lower concentrations. As the concentration gets very high, the relationship between absorbance and concentration can become non-linear. Essentially, the light path gets too "crowded" with absorbing molecules, and the simple proportionality breaks down. It's like trying to hear someone whisper in a really noisy room – eventually, you just can't make out what they're saying anymore, no matter how loud they try to whisper.
pH and Buffer Components: The ionization state of amino acids can be affected by pH. Changes in pH can subtly alter the absorption characteristics of the protein, leading to slight inaccuracies. Similarly, some buffer components might also absorb UV light, especially if they have aromatic structures.

Protein-Specific Absorbance (ε): The molar absorptivity (ε) is specific to each protein. While there are general estimates (like using an ε of 0.1 mL/(mg·cm) for a typical protein concentration of 1 mg/mL), if you know the exact sequence of your protein, you can calculate a more precise ε. This is where things get really accurate, but for many applications, a general value is good enough.
Making it Work for You: Calibration Curves and Known Proteins
So, how do we deal with these pesky real-world issues? We get smart! Two common strategies are:
1. The Calibration Curve (The Workhorse of Quantification)
This is probably the most common and robust method. Instead of relying on a theoretical ε, you create your own experimental relationship between absorbance and concentration using a known standard. Here's how it works:
- Prepare Standards: You take a protein of known concentration (a stock solution that's been accurately prepared, like Bovine Serum Albumin or BSA) and make a series of dilutions. You'll want several points covering a range of concentrations you expect your unknown samples to fall into.
- Measure Absorbance: You then measure the UV absorbance (at 280 nm) of each of these standards using your spectrophotometer and cuvettes.
- Plot the Graph: You plot a graph with concentration on the x-axis and absorbance on the y-axis. You should get a nice, straight line (or close to it!) – that's your calibration curve.
- Measure Your Unknown: Now, you measure the absorbance of your unknown protein sample(s).
- Interpolate: You find the absorbance of your unknown on the y-axis of your graph and then trace over to the line and down to the x-axis. The value on the x-axis is your protein's concentration!
This method accounts for the specific absorbance of your chosen standard protein and any consistent interfering substances present in your buffer system, making it much more reliable than just plugging values into the Beer-Lambert Law.
2. Using a Protein of Known Purity and Sequence
If you have a highly purified protein where you know its amino acid composition, you can often find or calculate its specific molar absorptivity (ε). For example, many databases provide this information. You can then use the Beer-Lambert Law (A = εbc) directly, plugging in your measured absorbance (A), the known path length (b), and the specific ε for your protein to calculate its concentration (c). This is great if you're working with a specific recombinant protein whose properties are well-characterized.

The Bradford and BCA Assays: When UV Isn't Enough
Now, while the UV spectrophotometry method is fast and non-destructive (meaning your protein isn't used up in the process), it's not always the best choice. What if your sample is heavily contaminated with nucleic acids? Or what if your protein is very low in tryptophan and tyrosine?
In these situations, scientists often turn to other colorimetric assays, like the Bradford assay or the BCA assay. These methods use chemical reactions that produce a colored product, the intensity of which is proportional to the protein concentration. You still use a spectrophotometer, but you're measuring absorbance at a different wavelength (usually in the visible light range) and you still need to create a calibration curve with a protein standard. The beauty of these methods is that they are generally less affected by nucleic acids and their sensitivity can be higher.
However, they are destructive assays, meaning your protein sample is consumed during the reaction. So, it's a trade-off! UV spectrophotometry remains a go-to for quick estimations, especially in applications where sample integrity is paramount.
Conclusion: A Handy Tool in Your Scientific Arsenal
So, there you have it. The humble UV spectrophotometer, with its ability to shine a specific beam of light through your protein soup and tell you roughly how much protein is in there, is an incredibly useful tool in any molecular biology or biochemistry lab. It’s not always the most precise method, and it has its limitations, but for a quick, dirty, and non-destructive estimate of protein concentration, it's hard to beat.
From figuring out if you successfully expressed your protein of interest to making sure you're adding the right amount to a reaction, this technology plays a crucial role. It’s a testament to how understanding the fundamental properties of molecules, like how they interact with light, can lead to powerful analytical techniques. So, the next time you’re staring at a cloudy solution and need to know what’s brewing, remember the UV spectrophotometer – your friendly neighborhood protein detective, always ready to shed some light (or rather, measure what's absorbed).
